Megakaryocyte Detection – EPFL & CHUV
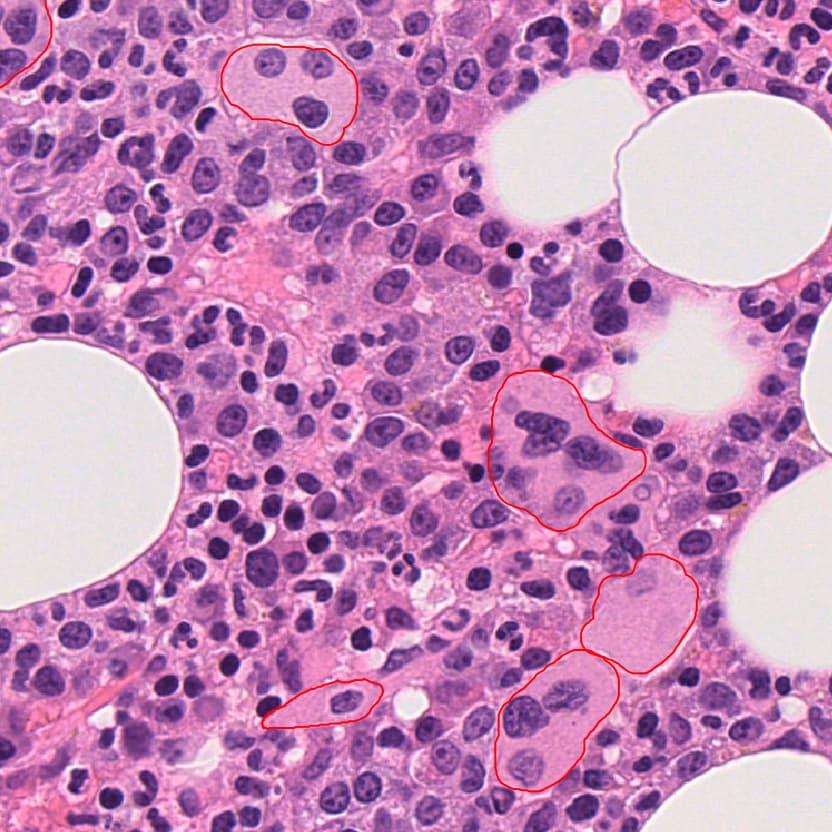
Built an automated pipeline for detecting and analyzing megakaryocytes (cells) in bone marrow histopathology slides, conducted as a Master’s semester project at EPFL in collaboration with CHUV.
The pipeline integrates deep-learning based segmentation (Cellpose for cell segmentation, Stardist for nuclear detection) with a custom ANN pixel classifier for bone structure segmentation.
Each detected cell is characterized by quantitative morphological and spatial features (area, nuclear lobulation, clustering via DBSCAN, distance to bone) implemented based on qualitative clinical tables.
The complete system was deployed as a QuPath extension featuring one-click execution, interactive feature visualization and display of extracted cell features, making the analysis accessible to non-programming clinical users.
Publication
Co-author of conference abstract presented at the Colloque Français d’Intelligence Artificielle en Imagerie Biomédicale (IABM’26), Lyon, March 9–11, 2026.
D. Sage, R. Sarkis, L.-F. Celma, M. Krause, J. Carretti, C.R. Chardon, N. Dewarrat, T. Smirnova, L. de Leval, O. Naveiras, “Towards Comprehensive Quantification of Morphological and Stromal Features in Bone Marrow Using AI-Driven Image Analysis”.